Enter a Search String |
| Special character and space not allowed in the query term. Search string should be at least 2 characters long. |
Parameters for CaRegulation (Pathway Number 132)
Pathway Layout
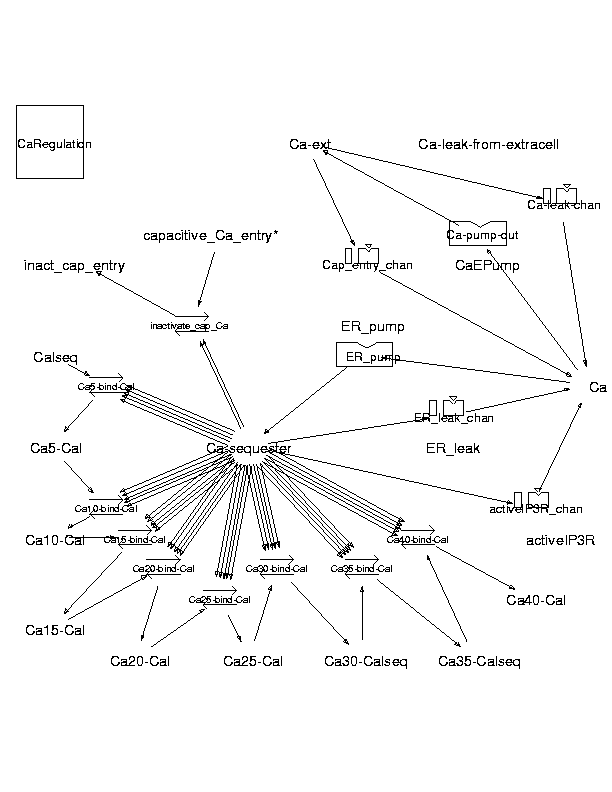
Pathway Basic Parameters
| Pathway Name | CaRegulation |
| Pathway Entry Number | 132 |
| Accession Number | 24 |
| Accession Name | Osc_Ca_IP3metabolism |
| Species | Generic mammalian |
| Tissue | Brain - Neuronal |
| CellCompartment | Cytosol |
| Related Pathways | 11, 86, 110, 149, 170 |
| Notes | Modified Ca Regulation model for the IP3 metabolism network. This model is used with the Othmer-Tang model for IP3 receptor kinetics, to generate cytosolic Ca oscillations. Channel kinetics of the IP3 receptor, ER-leak, ER-pump (or CaATPase) and Capacitative Ca entry have been modified to allow for Ca oscillations characterized in the Othmer-Tang model (Tang et al, Biophys J 70, 1996: 246-63). Kinetics of store Ca buffering by Calsequestrin have not been changed. |
Statistics
16 | 1 | 9 |
Conversion format
| This pathway is part of accession 24 and is completely specified in the file acc24.g. There is no separate files for just this pathway. |
| Format | File |
| Native Format (GENESIS format) | acc24.g |
| GENESIS Format (Annotated version) | Anno_acc24.g |
| Pathway Detail | Molecule List | Enzyme List | Reaction List |
